 
give a name and description of your list in the first 2 columns.leave the first 2 column emptys and paste using the special/ transpose option to paste your gene list horizontally on the Excel worksheet (first line).Open a new Excel workbook and transform the cell format to test (menu -> format -> cell -> text) make sure the gene identifiers of your list is the same as the ones used to create the enrichmentmap (e.g Gene Symbol).take the list of genes that you want to overlap with the map.How to create a signature gene-set to use with the post-analysis option of enrichmentmap We also can import attributes for these nodes. once the metanodes are created, we can change their size, color, shapes as other nodes.Run the "Annotate Cluster" with the option "Create Groups for clusters" checked (Control Panel -> Annotation Panel -> Advanced Option -> Create Groups for clusters). to expand a metanode, double-click on it and the individual nodes forming the group will be displayed.Īdd automatic labels using the "Annotate Cluster" feature (in menu -> EnrichmentMap -> Annotate Cluster).double click on the group on the node to collapse it.Right click on the selected nodes and choose the option "group" => "group selected nodes" => name it manual creation of groups and manual collapsing of nodes (1 by 1).Use cytoscape 3.2.1 or later and latest EnrichmentMap version (built 627 or later) How to collapse group nodes into a single metanode (in order to simplify the map) more on java folders and set JAVA_HOME environment variable to Java 7 path:. 
the recommended enrichmentmap version for Cytoscape 3.2 is the enrichmap version that contains the automatic annotation option (Release 2.1.0 ).You can use this script to upgrade your cytoscape 2.8 session files: convert_cyto_28_to_3_EM.sh or use the manual procedure described in the Cytoscape 3.1.1 section See instructions here: here if you have an older Snow Leopard 10.6.8 and need to upgrade to java 7 Once you installed java 7,open your terminal window and type "java -version" if the terminal tells you java version "1.6.x_xx", you may type this command "export JAVA_HOME=$(/usr/libexec/java_home -v 1.7)" to run java 7.To install java 7, you don't need to uninstall java 6 first.java7 is installed by default on (recent) Windows machines but not on MAC machines which run java 1.6 by default.It is based from received feedback from and tailored to the need of the OICR Cancer Stem Cell users ( )ĭo not hesitate to contact us if you need any assistance while executing these steps or have issues not reported on this page ( description on our workflow or how to create a maps, please visit these pages :, This page collects tips on how to use EnrichmentMap and Cytoscape based on frequently asked questions and known issues. 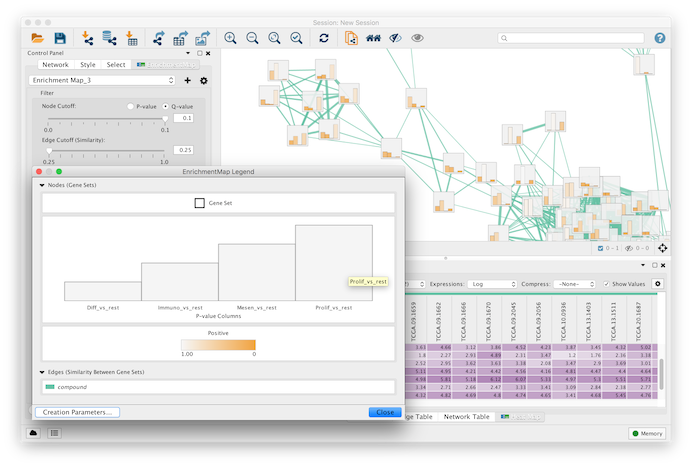
TIPS ON HOW TO USE ENRICHMENTMAP AND CYTOSCAPE (provided by the bioinformatics service and EnrichmentMap software developpers) How to make a Cytoscape 2.8 file compatible for Cytoscape 3 :.If you have a project done with Cytoscape 2.8 and want to correctly open these files with Cytoscape 3 you can follow this procedure (recommended for experienced users only)(tips created on Sept 23, 2014):.Move label position of nodes (Oct 5, 2014).Create a figure (manual annotation) with an EnrichmentMap in Cytoscape 3.1.1.Open an existing enrichment map session file (.cys) (Oct 5, 2014).Modify the node size to be proportional to GSEA NES (normalized enrichment score).How to create a signature gene-set to use with the post-analysis option of enrichmentmap.How to collapse group nodes into a single metanode (in order to simplify the map).TIPS ON HOW TO USE ENRICHMENTMAP AND CYTOSCAPE (provided by the bioinformatics service and EnrichmentMap software developpers).Style.url = paste(base.url, "styles", sep="/") Style <- list(title=style.name, defaults = defaults, mappings = mappings) Max.betweenness = max(V(graph1)$betweenness) Min.betweenness = min(V(graph1)$betweenness) VisualProperty="EDGE_TARGET_ARROW_SHAPE", Or, you can simply access the following URL from web browser: Here is the sample code to generate new Style: style.name = "R Style" # "value" : "DefaultVisualizableVisualProperty(id=EDGE, name=Edge Visu. # "visualProperty" : "DING_RENDERING_ENGINE_ROOT", = paste(base.url, "styles/default", sep="/") 
For example, if you want to see how default style is composed as JSON, you can get it by the following code. The best way to learn how to create them is getting preset Styles as JSON. You can create your own custom Visual Styles as R object. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
